Bifurcating FitzHugh-Nagumo
“I am a spec on a spec of a planet in the spec of a solar system in one galaxy of millions in the known Universe. My Universe is like a ring in the desert compared to the Footstool of God. And the Footstool like a ring in the desert compared to the Throne of God.”
–American Muslimah

Artist rendition of a neuron.
Attribution: HD Wallpapers
The amount of research done on brains, neurons, neuro-chemicals, etc….is astounding. No matter whom your favorite neuroscientist is you can rest assured that his or her knowledge barely scratches the surface of all that is known about human brains, or even Bumble Bee brains for that matter. But even all that’s known is but a “ring in the desert” compared to all that can be known–to all that will one day be known.
Hodgkin and Huxley developed their model of axonal behavior in the European Giant Squid in the late 1940s and published it in 1952. It was a game changer for two main reasons–first because it accounted for the action of Sodium and Potassium channels, and second because it stood up to experiment. It is a shining example of mathematical biological modeling from first principles.
FitzHugh and Nagumo based their model partially on the Hodgkin-Huxley equation, partly on the van der Pol equation, and partly on experimental data. While it’s not as accurate as the H-H equation relative to experiment, it’s much easier to analyze mathematically and captures most of the important attributes. It’s also been successfully applied to the behavior of neural ganglia and cardiac muscle. There are many different interpretations of the FitzHugh Nagumo systems of equations. The readiest on-line tool for examining them was created by Mike Martin.
In this post I’m working with a slightly modified version of the forms described by Ermentrout and Terman:
As you can see, this is a system of ordinary differential equations. The second equation has the quality of stabilizing the first. In a sense, it acts like an independent damper on changes in voltage when current is applied to an axon. Like the van der Pol equations, it’s non-linear and deterministic, and doesn’t easily lend itself to analytical solutions.
One thing you can’t see with Dr Martin’s tool is an interesting phenomenon that frequently occurs with these sort of equations: the Hopf Bifurcation. The only equation parameter that can be easily changed during experiment is the applied current. By programming the equations and calculating across a range of currents, it can be determined when an applied current produces unstable voltage oscillations. The point at which it becomes unstable is known as the critical point or Hopf point, and as long as the region of instability doesn’t extend to infinity, there will be one on each side. According to Steven Baer, the critical points for Ermentrout and Terman’s system are found at:
Enough about all that. Using parameter values of a = 0.8, epsilon = 0.5, and gamma = 0.2, I punched it out using my MatLab program FHNBifurc, and also ran it with XPP. XPP is no-frills and a little bit glitchy, but it’s awesome for solving systems of ODEs and it’s a champ of a program for bifurcation analysis. The installation is a little tricky, but it’s totally free–no spyware or anything else attached. If you haven’t at least toyed around with it, you should.
The MatLab program works by drawing the diagram in two directions–from the right in red and from the left in blue. The calculated critical points are shown as white stars. Ideally, the bifurcations should begin at those points, but as you can see, they don’t show up perfectly. That’s the image on the left. The best use of the program I’ve found is that it can be used to search across a wide range of currents to determine if and where instability occurs. I ran the same stuff with XPP’/Auto, and it gave me the figure on the right. Auto’s kind of a bear to get to the first time, but do it once and you’re set for life.
I also did a final run with Matlab, this time a smaller range with a lot more iterations. Took my computer about 10 minutes to complete it, so make sure you’ve got enough memory and processing speed before you try it. I edited-in the critical point solutions, FYI:
You can see why the FitzHugh-Nagumo equations are called eliptical. The MatLab program is great for scanning a wide range of values and locating the range of the bifurcation. Auto is way primo for drawing their shapes. Here’s the code for the XPP file:
# FitzHugh-Nagumo equations # fhnbifurc.ode dv/dt=-v*(v-1)*(v-a)-w+i dw/dt=e*(v-g*w) parameter a=0.8, e=0.5, g=0.2, i=0 @ total=50000, dt=1, xhi=50000., MAXSTOR=100000,meth=gear, tol=.01 done
The MatLab call looks like this:
FHNBifurc(a,eps,gamma,I1,IF,tspan,tstep)
a = value of a eps = value of epsilon gamma = value of gamma
I1 = initial current value IF = final current value
tspan = time span over which current is scanned
tstep = how short the time steps should be for the calculation
The specific call for the 0 to 10 scan was:
FHNBifurc(0.8,0.5,0.2,0,10,100000,0.01)
and for the more detailed diagram:
FHNBifurc(0.8,0.5,0.2,1.5,5,100000,0.001)
As always, the complete code is below. Cheers!
PS–Did you like The Sexy Universe on Facebook yet?
function FHNBifurc(a,eps,gamma,I1,IF,tspan,tstep)
steps = tspan/tstep;
Iramp = (IF-I1)/steps;
V(1) = 0;
W(1) = 0;
I(1) = I1;
for n = 1:(steps-1)
In = I(n);
Vn = V(n);
Wn = W(n);
fV = Vn*(Vn-1)*(Vn-a);
dVdt = -fV - Wn + In;
dWdt = eps*(Vn - gamma*Wn);
V(n+1) = Vn + dVdt*tstep;
W(n+1) = Wn + dWdt*tstep;
I(n+1) = In + Iramp;
end
IFwd = I;
VFwd = V;
WFwd = W;
V(1) = 0;
W(1) = 0;
I(1) = IF;
Iramp = -Iramp;
for n = 1:(steps-1)
In = I(n);
Vn = V(n);
Wn = W(n);
fV = Vn*(Vn-1)*(Vn-a);
dVdt = -fV - Wn + In;
dWdt = eps*(Vn - gamma*Wn);
V(n+1) = Vn + dVdt*tstep;
W(n+1) = Wn + dWdt*tstep;
I(n+1) = In + Iramp;
end
IRev = I;
VRev = V;
WRev = W;
Vcr(1) = (a+1-sqrt(a^2 - a + (1-3*eps*gamma)))/3;
Vcr(2) = (a+1+sqrt(a^2 - a + (1-3*eps*gamma)))/3;
for n=1:2
Icr(n) = (Vcr(n)/gamma) + Vcr(n)*(Vcr(n)-1)*(Vcr(n)-a);
end
Vcr
Icr
plot(IFwd,VFwd,'-r',IRev,VRev,'-b',Icr,Vcr,'*w')
xlabel('Current (I)')
ylabel('Voltage (V)')
Posted on December 29, 2012, in Bifurcation, Biology, MatLab, Neuroscience, Ordinary Differential Equations and tagged Bifurcation, Biology, MatLab, Neruoscience, Ordinary Differential Equations. Bookmark the permalink. Leave a comment.
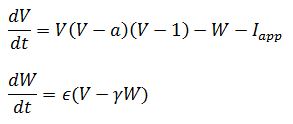
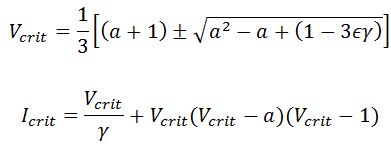


Leave a comment
Comments 0